Mass Photometry for Antibody–Antigen Binding and Stoichiometry
Quantify antibody–antigen binding, complex formation, and stoichiometry in minutes using nanograms of sample
Accurate, single-molecule, label-free analysis of antibody-antigen binding in solution
Mass photometry (MP) is transforming antibody characterization during development, enabling rapid, high-resolution analysis of antibodies and their interactions with antigens. MP directly detects individual molecules and complexes – including low-abundance species (<1%) – and allows rapid determination of antibody binding dissociation constants (KD). It delivers essential affinity and interaction data in significantly less time and using much less sample than conventional techniques.
Key pain points experienced by scientists developing antibody therapeutics:
Uncertain stoichiometry in complex systems:
For bispecific and multispecific antibody modalities, it is crucial to know the stoichiometry of complex formation as well as the affinity of each binding step, but this is not straightforward with conventional methods (e.g. biolayer interferometry (BLI), surface plasmon resonance (SPR)).
Challenges measuring at physiological concentrations:
Many analytical approaches require non-physiological conditions, reducing the relevance of the data for predicting in-vivo behavior.
Inconsistent antigen quality:
Variability in antigen integrity or activity can compromise binding studies, making rapid and reliable antigen QC an ongoing challenge.
High sample consumption:
Traditional techniques often consume large amounts of valuable material and require long assay times, which increases costs and limits throughput.
Challenges
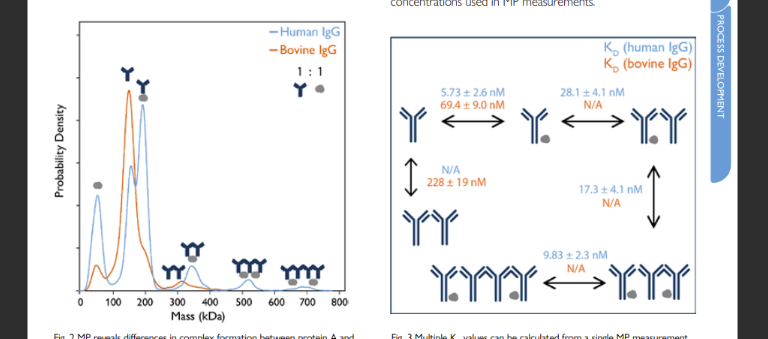
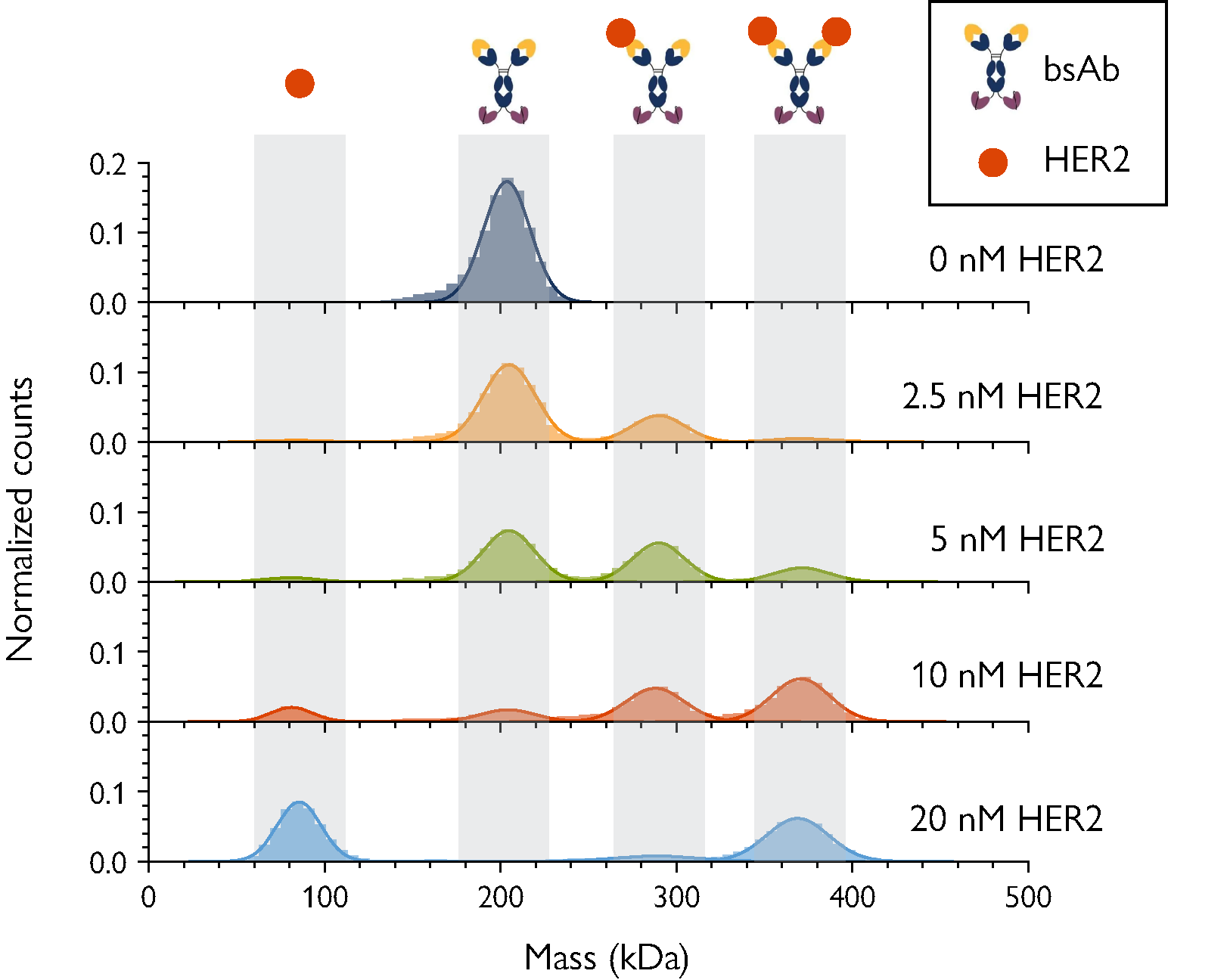
Figure 2. MP resolves complex bsAb-antigen interactions. The concentration of the antigen (HER2) was varied as indicated, with bsAb concentration fixed at 5 nM. As [HER2] increases, the peaks corresponding to free antigen as well as 2:1 Her2-bsAb complex become more prominent.
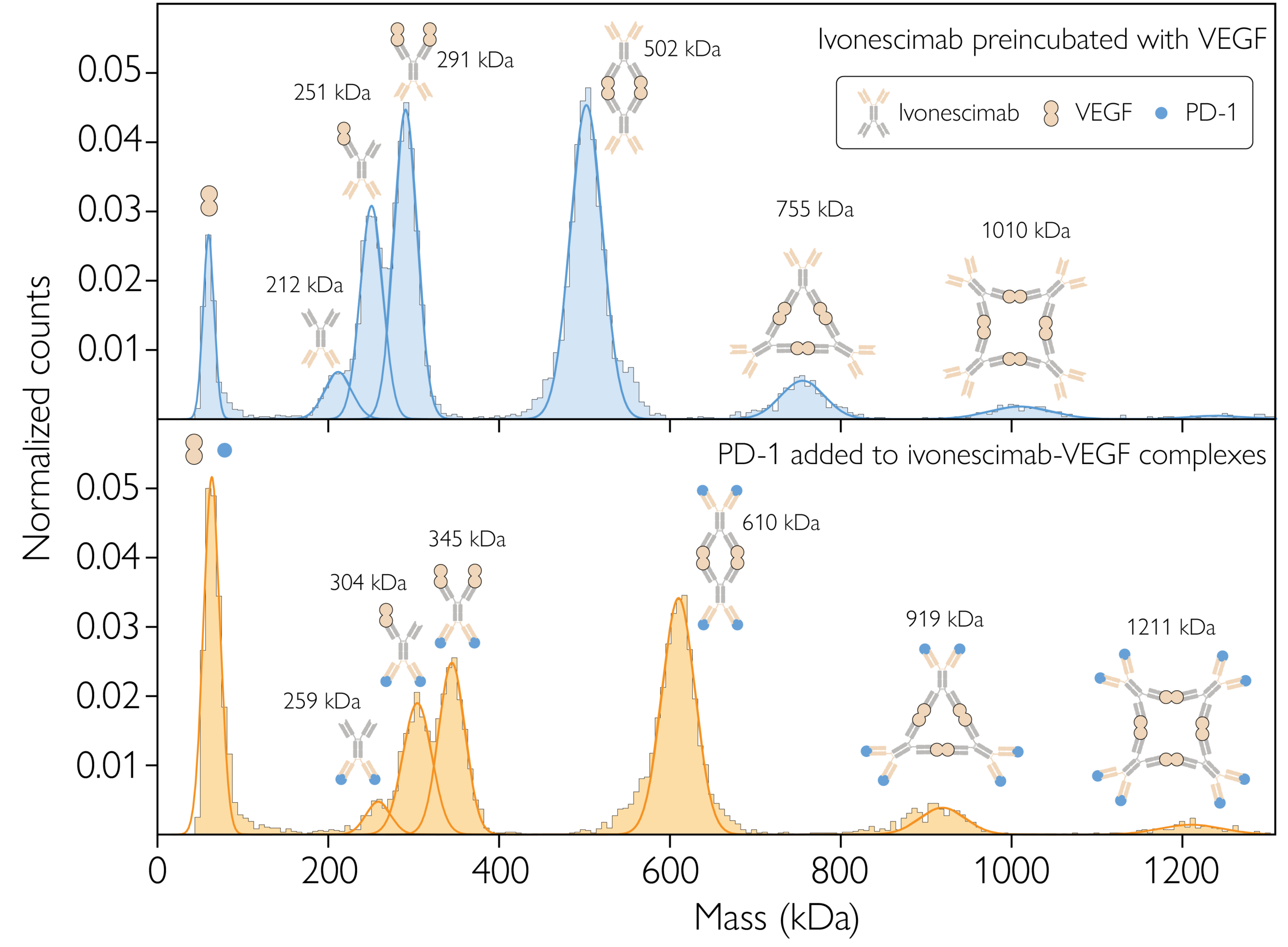
Figure 3. MP resolves the complexes formed from bispecific antibody ivonescimab interacting with its targets, VEGF and PD-1. Top: Ivonescimab–VEGF complex formation at equilibrium, with a 1:2 concentration ratio. Bottom: Ivonescimab–VEGF–PD-1 complex formation at equilibrium, with a 1:2:2 mixing ratio. Each plot shows the counts (normalized) of particles detected within each mass bin vs. the molecular mass. The schematics above each peak represent putative complex structures. Source: https://www.biorxiv.org/content/10.1101/2025.09.05.674400v2.full
Determining antibody binding stoichiometry and affinity
Efficient, reliable analytics are critical for assessing antibody affinity and binding behavior during development. A clear understanding of antibody-antigen interactions – including both stoichiometry and affinity – is essential for selecting and optimizing lead candidates.
Conventional techniques for assessing protein-protein interactions, such as BLI and SPR, provide valuable kinetic information – particularly for monovalent interactions. However, they rely on surface immobilization or labeling, which can affect binding activity. They also often require additional optimization and data fitting to accurately capture bivalent or multivalent interactions.
MP enables single-molecule analysis of antibody-antigen interactions in solution across a broad mass range (30 kDa – 5 MDa). As a fast, label-free technique requiring no immobilization, MP preserves native binding behavior and avoids the risk of obscuring binding sites – making it an ideal complement to SPR and BLI.
By directly detecting free species and complexes, MP reveals multiple antibody–antigen complexes in a single measurement, including stoichiometries beyond simple 1:1 or 1:2 binding. Quantitative equilibrium measurements allow dissociation constants (KD) to be determined directly from population counts. Low-affinity interactions (in the micromolar range) can be confirmed using Refeyn’s MassFluidix rapid dilution system. Multiple ligands or antigens can also be evaluated in parallel, accelerating comparative binding studies.
Physiologically relevant measurements
Although therapeutic antibodies are often administered at high concentrations (mg/mL or µM), their effective concentration at the target site is typically in the nanomolar range. Understanding antibody-antigen interactions under these physiologically relevant conditions is therefore critical for predicting in vivo efficacy and guiding development decisions.
MP provides unmatched sensitivity for studying binding in real time at nanomolar concentrations, label-free and in solution. Unlike surface-based techniques such as SPR and BLI – where immobilization can restrict native complex formation, MP measures interactions in a soluble environment. This preserves natural binding stoichiometries and enables the detection of multiple antibody-antigen assemblies in a single measurement.
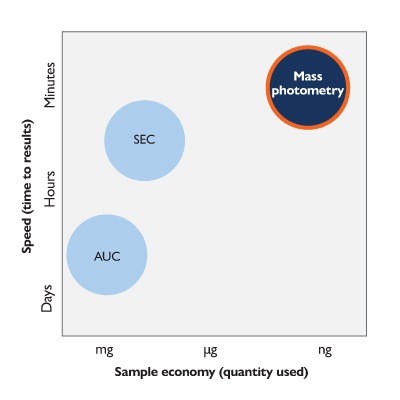
In addition, conventional in-vitro methods such as size-exclusion chromatography (SEC) and analytical ultracentrifugation (AUC), when performed without signal amplification (e.g. fluorescent probes), cannot reliably detect protein complexes at low nanomolar concentrations due to limitations in sensitivity. MP overcomes these constraints, enabling direct observation of immune complex formation under physiologically relevant concentrations.
MP – combined with the MassFluidix rapid dilution system – allows researchers to study high- and low-affinity interactions in conditions that closely mimic the native environment, accelerating development and improving confidence in candidate selection.
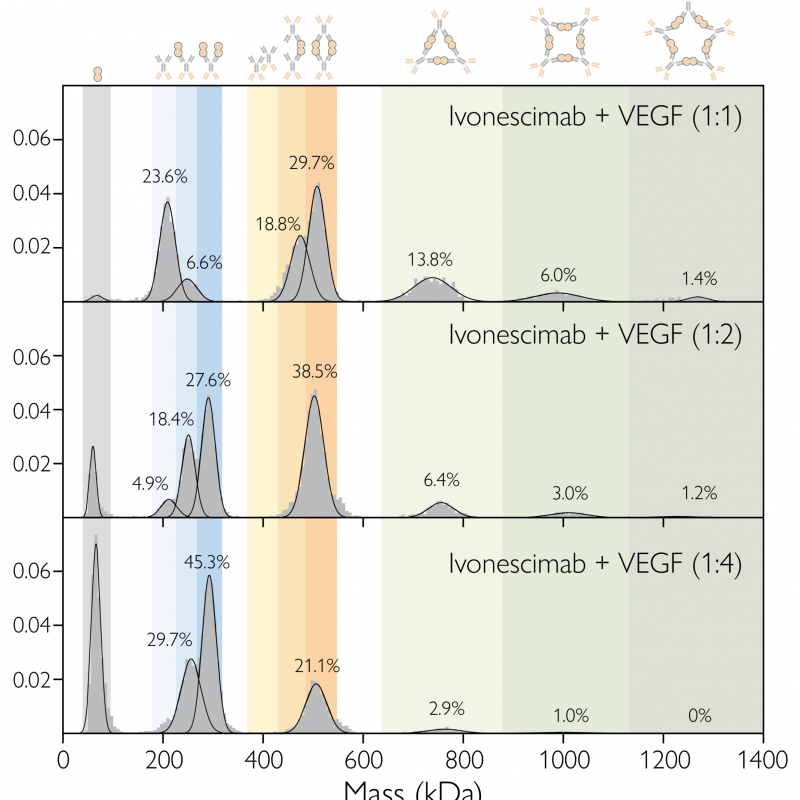
Figure 4. Probing different antibody antigen molar ratios at nM concentrations? Ivonescimab:VEGF complex formation for mixtures with 1:1, 1:2 and 1:4 antibody: antigen concentration ratios, at equilibrium, after 40 min incubation, at nM concentration.
Ensuring antigen quality prior to kinetic analysis
The reliability of antibody binding data depends not only on the analytical method, but also on the quality and behavior of the antigen. Before progressing to kinetic analysis with surface-based techniques, such as SPR or BLI, it is essential to confirm that both ligand and analytics are in their preferred oligomeric states and behave as expected in solution.
Changes in antigen or analyte oligomerization can significantly distort binding measurements. For example, if an analyte partially oligomerizes when it should remain monomeric, apparent binding strength may be overestimated – leading to incorrect conclusions about affinity and mechanism of action. Without early verification, such effects could go unnoticed and compromise decision-making downstream.
MP enables rapid, label-free assessment of antigen quality prior to SPR or BLI experiments. By directly measuring and reporting species populations in solution, MP can confirm whether antigens and antibodies are present in the expected oligomeric form. It can also confirm the stoichiometry and affinity of the complexes they form. In this way, MP provides rapid, essential reassurance before committing samples to surface-based kinetic assays.
MP also allows fast screening of buffer and formulation conditions, revealing how these conditions affect antibody and antigen behavior. This capability supports rapid optimization of assay conditions, ensuring the SPR and BLI experiments are performed with suitable reagents – and supporting meaningful interpretation of the kinetic data obtained.
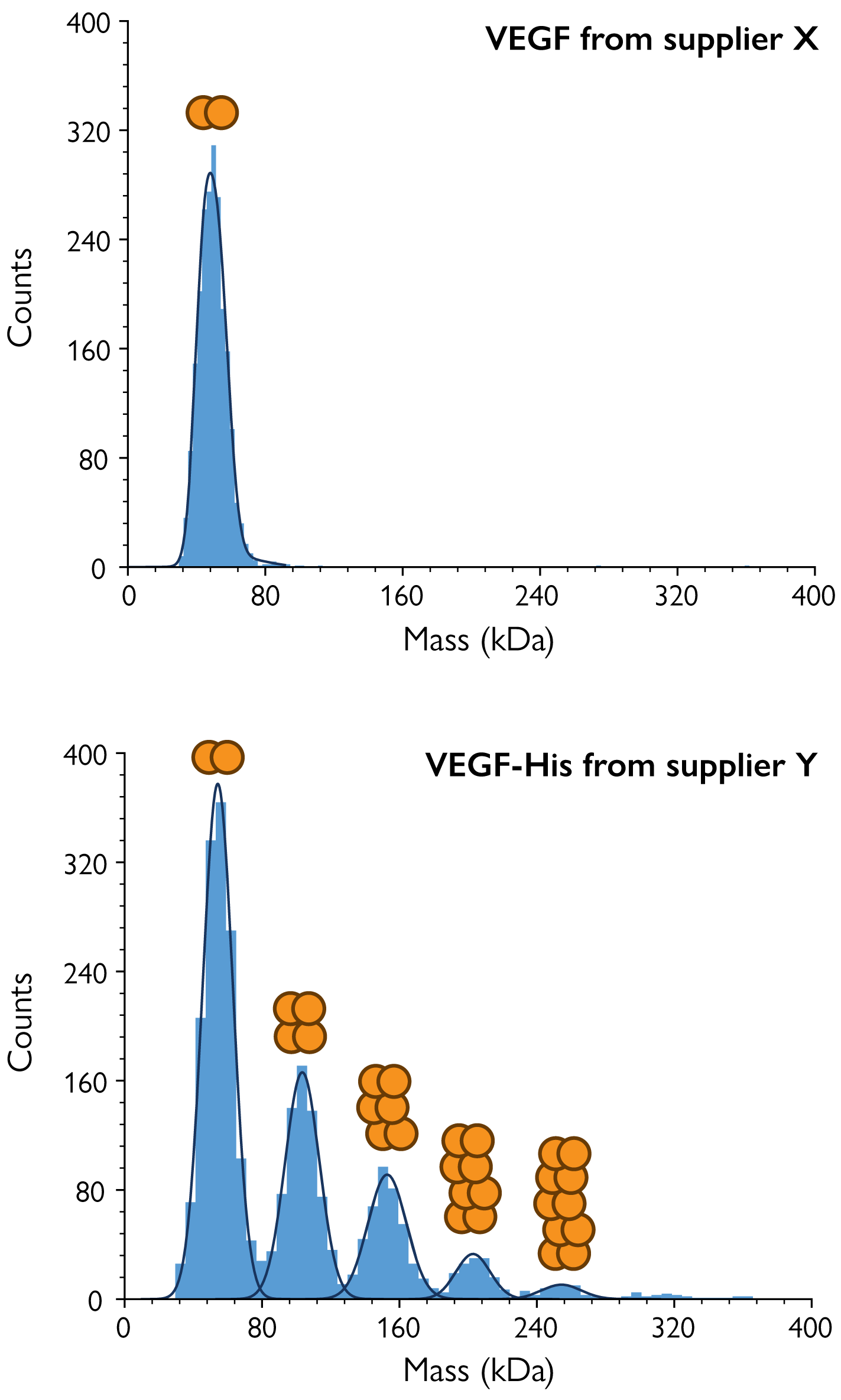
Figure 5. MP reveals differences in VEGF oligomerization between suppliers. MP was used to measure samples of the antigen VEGF from supplier X (top) and VEGF-His tag from supplier Y (bottom). MP clearly showed significant oligomerization in the supplier Y sample, although VEGF naturally exists as a dimer. Both samples were tested in PBS buffer at nanomolar concentrations.
Conventional techniques consume too much sample
Analytical ultracentrifugation (AUC) and size-exclusion chromatography (SEC) are commonly used to characterize protein complexes, determine oligomeric states, and assess antibody–antigen interactions. However, these methods typically require microgram-to-milligram quantities of material – making frequent analysis unfeasible and driving up development costs.
MP overcomes this limitation by providing quantitative binding, stoichiometry, and sample quality data using just nanograms of protein. It enables rapid assessment of interactions, evaluation of complex formation, and buffer optimization – all with just a droplet of sample.
Figure 5. Mass photometry takes less than 5 minutes and uses very little sample (just tens of µL at nanomolar concentration). This makes it ideal not only for antibody-antigen binding studies, but also for fast screening of buffer and formulation conditions and checking reagent quality and oligomeric state before progressing to SPR or BLI.
Mass photometry solutions for antibody analysis
TwoMP / TwoMP Auto
The TwoMP provides unique insight into antibody samples by measuring the mass of individual molecules in native conditions – completely offering label-free and with single-molecule resolution.

It enables accurate, multi-attribute characterization of antibody therapeutics, including complex modalities, with minimal sample preparation and nanograms of sample. Designed for intuitive operation, it delivers reliable results within minutes.
The TwoMP Auto extends these capabilities with automated handling for up to 24 measurements – increasing throughput, improving repeatability, and offering walkaway timeIt delivers unique insights into LVV preparations by simultaneously measuring particle size and contrast (a proxy for mass) – enabling rapid assessment of vector purity, composition, and yield. This makes it an ideal tool for fast, label-free analysis.
Its core features address key analytical challenges in antibody aggregation:
Minimal Sample Requirements
Requires only nanograms of sample for analysis, preserving precious material.
Rapid Analysis
Delivers comprehensive antibody aggregation data within minutes per sample.
Native, Single-Molecule Measurement
Measures molecular mass with single-molecule resolution and under native conditions.
MassFluidix™ HC System
Refeyn’s microfluidics system, MassFluidix HC, raises the upper sample concentration limit from the nanomolar to the micromolar range.

Significantly expanding the sample concentration range amenable to investigation by mass photometry, MassFluidix HC is an add-on for Refeyn’s TwoMP mass photometer. It includes a central unit with fluid controls. Ready-to-use rapid-dilution microfluidics chips are also available from Refeyn for this system.
MassFluidix offers:
Characterization of concentrated samples
Starting concentration up to the tens of micromolar.
Rapid analysis requiring little sample
~20 min for setup and cleaning, using 20–60 µL of sample.
User-friendly operation
User training takes just one day, supported by intuitive fluid-control software.
MassGlass™ UC Slides
High-quality mass photometry measurements begin with clean slides, and MassGlass UC provides a pristine surface, straight out of the box.
Refeyn offers MassGlass UC – Sample Prep Kits for the TwoMP and TwoMP Auto systems, containing everything you need for a seamless experience when measuring your antibody samples.
MassFerence™ P1 Calibrant
This protein calibration standard is essential for mass photometry measurements, offering maximum precision with minimum hassle.
The MassFerence P1 protein standard is recommended for use with the TwoMP and TwoMP Auto, and is suitable to use when measuring samples containing proteins with mass ranging from 90 to 1000 kDa.
Mass photometry in antibody binding and stoichiometry literature
Resources

Rapid analysis of bispecific antibody stability and target binding by mass photometry
Analyzing bispecific antibodies can be challenging, as commonly used techniques struggle to provide information on their different binding sites. This application note shows how mass photometry efficiently characterizes the purity, stability, and binding of multiple bispecific antibody candidates.

Explore the binding mechanism of bispecific antibody ivonescimab with mass photometry
In this webinar, we will show how we used mass photometry, a single-molecule imaging technique capable of detecting biomolecular complexes in solution, to characterize ivonescimab’s binding behavior with VEGF and PD-1.

Characterizing Antibody Binding to SARS-CoV-2 with Mass Photometry
Characterizing the mass of proteins and their complexes is essential for understanding biomolecular interactions. However, heterogeneity in biological systems has made this a challenge for traditional native mass spectrometry and structural biology techniques. In this webinar, Victor Yin (Utrecht University) discusses how mass photometry and charge detection mass spectrometry (CDMS) enable the study of complex protein systems, including full antibody interactions with the SARS-CoV-2 spike protein.

Mass photometry reveals stoichiometry and binding dynamics of bispecific tetravalent anti-VEGF-PD-1 antibody ivonescimab
MP enabled detailed analysis of bispecific antibody Ivonescimab complex stoichiometry interactions with VEGF and PD1.
MP confirmed VEGF-induced ivonescimab oligomerization
Results show dimers, not higher-order structures, were the most stable stoichiometry for Ivonescimab-VEGF
Affinity constants were determined for each ivonesicmab-VEGF complex stoichiometry
Kinetic parameters for PD-1 interaction with Ivonescimab were calculated
Frequently asked questions
What types of antibody–antigen interactions can mass photometry measure?
Mass photometry can measure interactions, as long as the mass of each molecule is above 30 kDa, there is a resolvable difference in mass between the antigen and antibody, and the interaction occurs at nanomolar concentrations (for mass photometry alone) or up to the tens of micromolar concentrations (for mass photometry with the rapid-dilution MassFluidix system).
In terms of resolution, for mass photometry, resolution varies across the mass range. It is 25 kDa at 66 kDa, for instance –
learn more in our article, Understanding the strengths and limitations of mass photometry.
How can mass photometry be used to determine antibody binding stoichiometry?
Mass photometry measures individual proteins and complexes directly in solution, without relying on labels or surfaces. Unlike traditional methods that rely on UV absorbance – where larger complexes can skew the signal – MP detects every particle individually. This allows accurate determination of antibody–antigen stoichiometry, showing how many antigens bind to each antibody under native conditions.
Can mass photometry differentiate between monomeric, dimeric, and higher-order antibody aggregates?
Yes, it can differentiate between antibody aggregates and it can also characterize the effect of aggregates on binding and stoichiometry. Visit our page on measuring antibody aggregation with mass photometry to learn more.
What sample concentrations are required for accurate antibody analysis with mass photometry?
Mass photometry can measure antibody-antigen complexes formed at nanomolar concentrations, making it ideal for studying interactions at physiologically relevant levels. For interactions that occur at micromolar concentrations, mass photometry can still be used – by applying the MassFluidix rapid dilution microfluidics system, which brings the sample into the optimal measurement range for measurement without disrupting complex formation. This flexibility allows MP to analyze a wide range of binding affinities and complex stoichiometries.
Can mass photometry be used to analyze antibody–antigen binding dissociation constants (Kd)
Yes, as demonstrated in Kofinova et al (2024) and Wu and Piszczek (2020), mass photometry can be used to determine the mass distributions of binding partners, the complexes they form, the relative abundance of each species, and, accordingly, the dissociation constant (KD) of their interactions.
Can mass photometry analyze bispecific or multispecific antibody binding and complex formation?
Yes, as mass photometry has high resolution and sensitivity, it can easily identify multiple antibody-antigen complexes. This has been demonstrated, for example, in Jajcanin Jozic et al (2026).
How does mass photometry integrate with other analytical techniques in antibody characterization workflows?
During early R&D and discovery stages, MP complements traditional binding studies by rapidly assessing reagent quality and antibody-antigen stoichiometry (in just minutes using nanograms of sample). It provides confidence in the reagents before investing time and resources into more complex techniques, like SPR.
At later stages in Development, MP is a column-free orthogonal to SEC for aggregation assessment when comparing different candidates or conditions e.g. in forced degradation / accelerated stability studies. MP is 300x faster and uses at least 8000x less sample than AUC – the current technique used as a column-free orthogonal to SEC.
How does MP measure affinity constants and is that a feature in the software?
The approach is described in detail in Kofinova et al (2024), but this is currently not feature in the software.
How does mass photometry work?
To learn more about how mass photometry works, we recommend our article, How does mass photometry work?; the webinar, Measuring molecules with light; and our e-book, Understanding mass photometry.
Discover how mass photometry quantifies antibody – antigen binding in minutes
Ready to revolutionize your research?
Find out how mass photometry can transform your sample characterization. Whether you’re focused on drug development, discovery science, or manufacturing, Refeyn’s innovative technology provides the insights you need to accelerate your breakthroughs.
If you have any other questions or would like further details, please get in touch!