Sample purity and optimization
Structural studies of protein assemblies depend on good sample quality. Mass photometry supports every step – from early sample screening to pre-cryo-EM quality control – using only nanograms of sample
See every protein and complex between 30 kDa and 5 MDa in minutes with single-molecule resolution.
Successful structural studies require samples that are homogeneous, intact, and stable. Before committing time and resources to cryo-EM or native MS, structural biologists need confidence that their sample is ready – but the conventional toolkit of SEC, DLS, SDS-PAGE, and negative stain EM has significant limitations.
Key pain points experienced by researchers:
Missed heterogeneity:
Ensemble techniques can fail to detect sample polydispersity, leading to failed structural experiments.
Speed vs. resolution trade-off:
Most techniques with single-molecule resolution are too slow for routine condition screening.
Limited sample:
Characterization and optimization consume too much precious material.
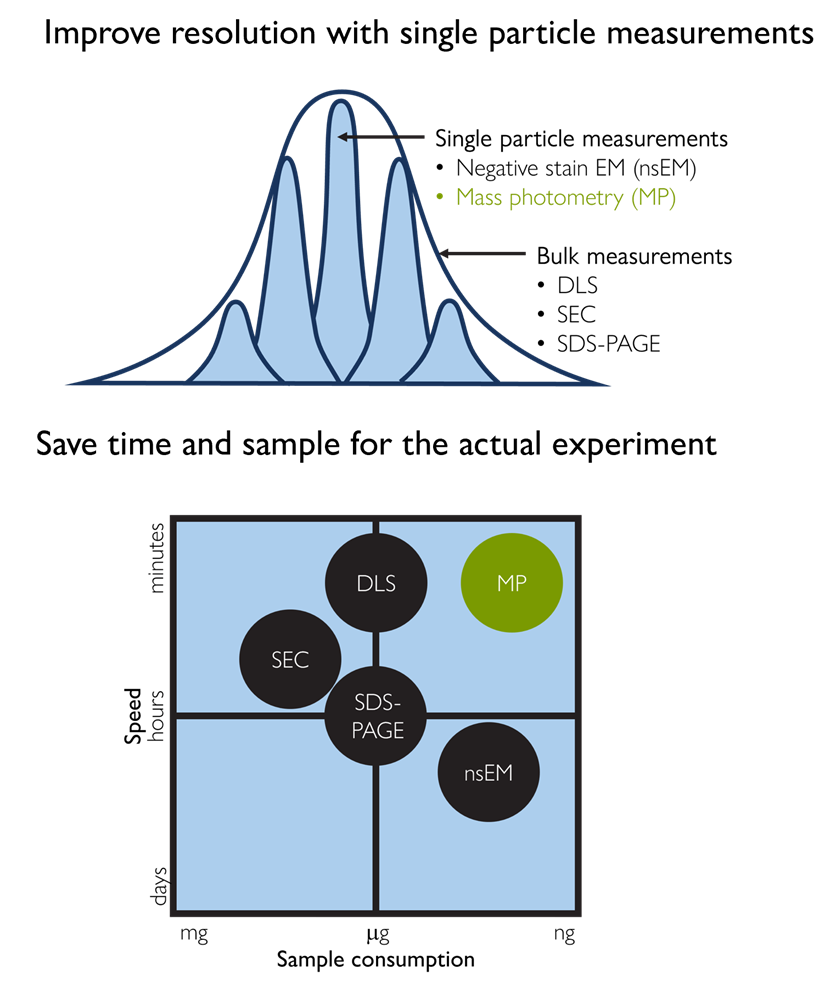
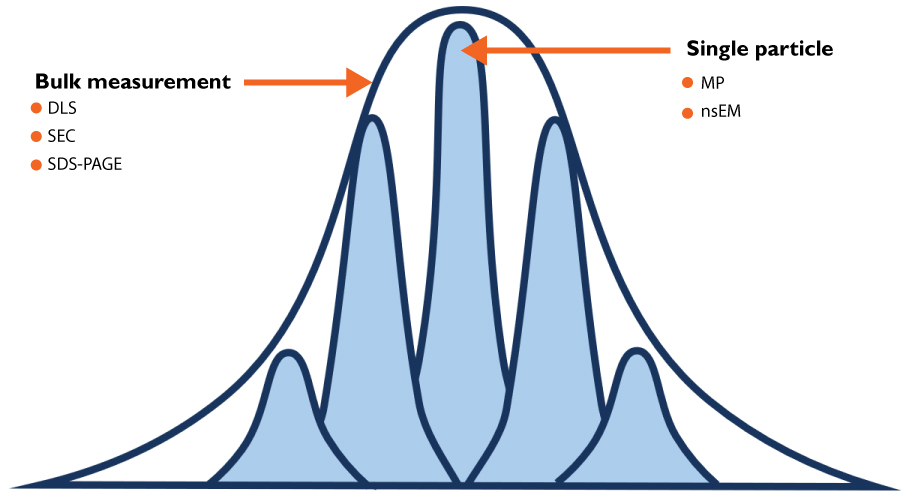
Figure 1. Mass photometry combines speed, resolution, and low sample consumption. Conventional ensemble techniques, such as SEC, DLS, and SDS-PAGE, are fast but lack single-molecule resolution. Negative stain EM (nsEM) offers that resolution but is slow and resource-intensive. Mass photometry occupies a unique position, delivering single-molecule resolution with rapid turnaround and minimal sample consumption.
Challenges
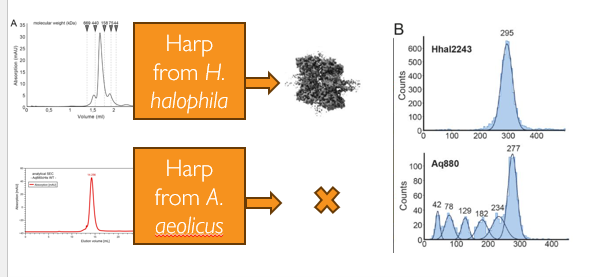
Figure 2. Mass photometry reveals hidden polydispersity missed by SEC. Despite similar SEC profiles, mass photometry (MP) showed that for one HARP oligomer (Aq880), the main species, made up only half the sample, with multiple LMWS also present. (Adapted from Feyh et al., eLife, 2021)
SEC, DLS, and SDS-PAGE can miss critical sample heterogeneity
High-quality structural analysis starts with a high-quality sample. But conventional, bulk techniques – such as SEC, DLS, and SDS-PAGE – report on the average behavior of a population, and can fail to resolve subtle differences in composition or oligomeric state that would be apparent from single-molecule measurements.
When sample heterogeneity goes undetected, the consequences are significant: Failed structural experiments, wasted reagents, and large losses of money and cryo-EM or native MS time.
Because mass photometry measures the mass of individual molecules in solution, it reveals the true composition of a sample. Where ensemble techniques may show a single, apparently homogeneous peak, mass photometry resolves heterogeneity. It provides the resolution needed to have confidence in your sample before committing to downstream analysis.
Optimizing conditions for structural studies requires speed and single-molecule resolution
Screening buffer conditions – pH, salt concentration, crosslinker ratios – is a routine but time-consuming part of sample preparation for structural studies. Negative stain EM can provide single-molecule resolution on sample heterogeneity, but is slow, resource-intensive, and requires special expertise.
Mass photometry provides a clear readout of sample purity and the integrity of complexes, and it works in minutes using nanograms of sample. It is compatible with a wide range of buffer conditions, making it well suited to rapid condition screening. Mass photometry measurements are accurate and agree with nsEM results – but can be obtained in a fraction of the time.
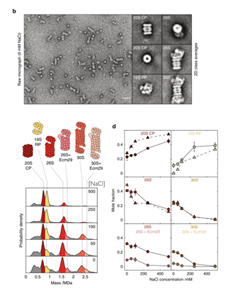
Figure 3. Mass photometry enables rapid screening of buffer conditions for proteasome stability and agrees with nsEM. Mass photometry (MP) histograms show the 30S proteasome dissociating into smaller subunits at increasing NaCl concentrations, with the relative abundance of each species consistent between the two techniques. (Adapted from Sonn-Segev et al., Nature Communications, 2020)
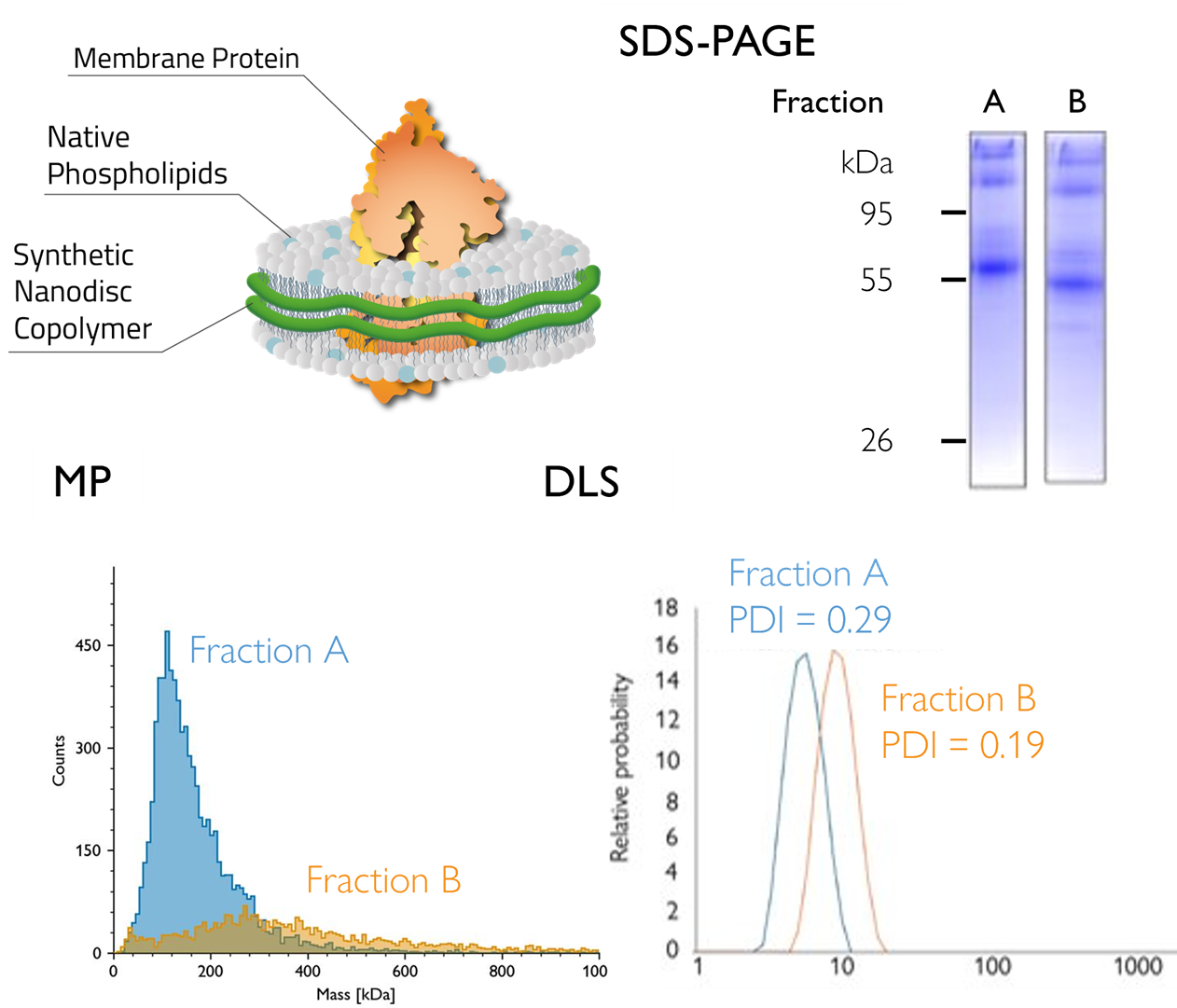
Figure 4: Mass photometry identifies the optimal SEC fraction of a nanodisc-embedded GPCR. MP analysis showed one fraction of GPCR GLP1 (solubilized in nanodiscs) was more homogeneous, with a mass consistent with the expected value (~110 kDa) – information not resolvable by dynamic light scattering (DLS) alone. Data obtained in partnership with Cube Biotech. (Adapted from Mass photometry analysis of GPCRs in copolymer nanodiscs)
Reserve enough sample for downstream analysis
Many proteins of structural interest are difficult or expensive to produce in large quantities – particularly those requiring eukaryotic expression systems, primary cells or tissues, or incorporation into membrane mimetics such as nanodiscs. When sample is scarce, every microliter spent on characterization is a microliter unavailable for functional or structural studies.
Mass photometry delivers a direct assessment of sample homogeneity and mass distribution using nanograms of material in minutes. Its low sample consumption makes it well suited to structure-function workflows where sample is limited. It enables thorough characterization while leaving sufficient sample for downstream analysis.
Mass photometry solutions for sample quality assessment and optimization
Refeyn mass photometers measure the mass of individual molecules in solution, delivering a clear readout of sample purity, homogeneity, and complex integrity in minutes – using just nanograms of material. Compatible with a wide range of buffers, they are well suited to rapid sample checks and screening buffer conditions for structural biology and biophysics workflows.
MyMass

MyMass™ is a compact, intuitive mass photometer designed for rapid, label-free characterization of proteins and complexes at the single-molecule level. Built for structural biologists and others who need a fast, reliable sample quality check before committing to more complex, costly analysis.
Refeyn TwoMP & TwoMP Auto

TwoMP, Refeyn’s flagship mass photometer, is a powerful instrument offering the highest performance for demanding structure and function investigations. TwoMP Auto adds automated sample handling for consistent, walkaway measurements across multiple samples.
Two optional add-ons for TwoMP extend its capabilities for more demanding structural biology or biophysical workflows.
MassFluidix™ HC System

Standard mass photometry requires samples to be diluted to the nanomolar range for measurement. The MassFluidix HC microfluidics system removes this limitation by rapidly diluting samples from starting concentrations of up to tens of micromolar directly on-chip – capturing the sample’s native state before dilution-driven dissociation occurs. This makes it particularly valuable for studying weak or transient interactions, such as low-affinity protein complexes, that would otherwise be undetectable.
TwoMP Auto
An automation unit adds automated sample handling to the TwoMP, enabling consistent, walkaway measurements across multiple samples. TwoMP Auto can autonomously analyze 24 samples in as little as 90 minutes – ideal for labs screening multiple buffer conditions or protein variants in parallel.
Refeyn consumables are engineered for mass photometry, to ensure reliable, high-quality measurements every time. The offering includes:
MassGlass™ UC
MassGlass UC slides are high-quality measurement surfaces designed for mass photometry analysis of proteins. They provide an out-of-the-box, ready-to-use solution that delivers reliable and consistent performance, ensuring high-quality data with every measurement. Compatible with MyMass, TwoMP, and TwoMP Auto.
MassFerence™ P1 Calibrant
MassFerence P1 is a calibration standard designed to ensure accurate, reproducible mass photometry measurements across experiments, instruments, and laboratories. For use when measuring proteins spanning 90 to 1000 kDa, providing reliable calibration across a broad molecular mass range. Compatible with MyMass, TwoMP, and TwoMP Auto.
For a full range of consumables and kits for all your mass photometry need, visit the Refeyn eShop.
MyMass integrated software
MyMass features built-in software designed to be intuitive and accessible, even for occasional users. Guided workflows with diagrams and videos at every step walk users through data acquisition, while extensive help documentation and FAQs provide further support.
The software also makes analysis straightforward – mass distributions are displayed with automatic peak fitting, adjustable axes, and a clear table of fitted peak masses and relative abundances. Data can be exported as a PNG of the mass distribution or a CSV of the fitted peak table, making it simple to incorporate results into lab notebooks and reports.
AcquireMP & DiscoverMP
For TwoMP and TwoMP Auto users, Refeyn’s AcquireMP and DiscoverMP software suite provides a complete solution for data acquisition and analysis. AcquireMP is the primary software used to control the instrument, and allows users to record movies of their measurement, and adjust and optimize parameters for data acquisition. DiscoverMP, the data analysis package, makes it straightforward to inspect and interpret datasets, and generate and export figures.
Mass photometry for sample quality assessment – peer-reviewed evidence
Quantifying the heterogeneity of macromolecular machines by mass photometry.
Sonn-Segev et al benchmark MP against negative stain EM, SDS-PAGE, and SEC for assessing the heterogeneity, integrity, and assembly state of large complexes including the proteasome and APC/C – highlighting its value for sample screening and workflow optimization.
Need for speed: Mass photometry as a sample analysis tool for structural studies
Discusses the principles of mass photometry and its application as a valuable tool for structural biologists – demonstrating how it streamlines cryo-EM pipelines by providing rapid, label-free sample quality assessment at the single-molecule level.
Higher-order structure and proteoforms of co-occurring C4b-binding protein assemblies in human serum
Kadavá et al employ MP together with charge-detection MS, crosslinking MS, and high-speed atomic force microscopy to resolve and quantify co-occurring isoforms of C4BP. This integrative structural approach expands the understanding of complement regulation and the dynamic nature of C4BP complexes in human serum.
Structure and mechanistic features of the prokaryotic minimal RNase P
Feyh et al use MP to investigate the oligomerization behavior of a minimal RNAse P system, correlating its oligomeric state with enzyme activity and using MP to screen variants with more uniform oligomeric states to increase the likelihood of obtaining a structure.
Mass Photometry of Membrane Proteins
Olerinyova et al employ mass photometry to characterize membrane proteins in a range of membrane mimetic systems, such as detergents, amphipols, nanodiscs. They demonstrate that particle size, sample purity, and heterogeneity can be accurately determined to support high-resolution structural studies of membrane proteins.
Frequently asked questions
How does mass photometry work?
Mass photometry is an analytical method that measures molecular mass by quantifying light scattering from individual biomolecules and particles in solution. It measures the interference between the light scattered by the molecule and the light reflected by the measurement surface. The signal measured is called the contrast (or interferometric contrast) and is directly correlated with molecular mass, as demonstrated by the seminal paper introducing mass photometry (Young et al., 2018).
To learn more, we recommend the following resources:
How much sample is needed for a measurement?
Very little sample is needed.
- For MyMass, the requirement is 30 µL of sample at a concentration of 1–100 nM (10 nM optimal).
- For TwoMP (or TwoMP Auto), we recommend starting with 20 µL of sample at a concentration of 10 nM, adjusting if needed. This corresponds to 10 nanograms of a 50 kDa protein or 100 nanograms of a 500 kDa protein.
How can mass photometry be used to screen for sample heterogeneity?
Mass photometry screens for sample heterogeneity by directly measuring the mass of individual molecules in solution, eliminating ensemble averaging artifacts and revealing whether a sample contains a single, well defined species or a mixture of subassemblies and impurities. Because MP operates under native, non denaturing conditions and is insensitive to molecular shape, it can accurately assess the mass and stability of non covalent complexes – even when they are structurally flexible – providing a rapid and detailed view of sample homogeneity.
How can mass photometry (MP) be used to determine the oligomeric state of a protein?
Mass photometry measures the mass of individual molecules in solution, independent of their shape, and is compatible with native buffers. The near-native, single-molecule mass readout informs on the stoichiometry of non-covalent interactions and complexes across a broad mass range. It also has a broad dynamic range, capable of detecting very rare and very abundant species.
How can I conserve my sample when screening buffer conditions?
Mass photometry allows users to optimize buffer conditions that favor protein complex stability, uniformity, and desired stoichiometry using nanograms of sample and small volumes of reagents in minutes. Mass photometry is compatible with a variety of buffers and requires little to no method development, allowing the effects of altering pH, salt concentration, small molecules, etc. on sample homogeneity to be easily evaluated.
What is the resolution of mass photometry measurements?
The mass resolution of mass photometry measurements depends on the sample’s mass, heterogeneity, and overall quality, and is therefore best determined experimentally for each case.
In addition, a way to estimate the minimal mass difference that can be distinguished between two species is to examine the sigma (σ) of the Gaussian fit for a population in the mass distribution. Multiplying σ by 2.35 yields the full width at half maximum (FWHM), which corresponds to the smallest mass difference that can be resolved between two species of similar mass.
Resources
Article: Enhancing cryo-EM workflows with mass photometry: A smarter pre-screening approach
Discover how mass photometry can streamline your cryo-EM pipeline – from early sample screening to pre-submission quality checks. It saves time and reduces the risk of failed cryo-EM analysis.
Webinar: Structural characterization of blood biomolecular assemblies with mass photometry
Hear from Prof. Dr. Albert J. R. Heck, on how mass photometry can characterize complex, co-occurring protein assemblies in our blood – and how it complements orthogonal structural biology techniques, such as (glyco)proteomics, cross-linking mass spectrometry (MS), native MS, computational modelling and single-molecule MS.

Rapid, high-resolution protein characterization with MyMassMP
Discover how MyMassMP quickly assesses sample purity and homogeneity across a broad mass range – with just nanograms of protein and results in one minute. An alternative to SDS-PAGE, SEC, and DLS for routine sample evaluation.

Quantifying macromolecular heterogeneity of macromolecular machines by mass photometry
Hear Dr Adar Sonn-Segev describe how mass photometry provides quantitative information on samplecheterogeneityy using minimum volumes with molecular resolution.
Application Note: Characterization of protein interaction equilibria
See how MP makes it possible to monitor complex equilibrium formation reactions, and to assess how changes in the chemical environment or protein concentration affect equilibria.
Application Note: Characterizing protein oligomerization with automated mass photometry
Learn how MP can measure the mass distribution of a sample in native conditions and at a single-molecule level, with enough sensitivity to detect rare species. Automated mass photometry builds on these capabilities with easy and consistent sample dilution and manipulation, enabling thorough assays of oligomerization behavior.
Learn how mass photometry can accelerate your sample assessment and optimization
Ready to revolutionize your research?
Find out how mass photometry can transform your sample characterization. Whether you’re focused on drug development, discovery science, or manufacturing, Refeyn’s innovative technology provides the insights you need to accelerate your breakthroughs.
If you have any other questions or would like further details, please get in touch!